Multicolor stack in imagej fiji3/14/2024 It was mostly intended to help with newer 7-8+ channel fluorescent images:įinally, for percentage area, I also use a script that sums the areas of objects of a particular set of classes, and then divides them by the total area of their parent annotation, and adds that percentage to the parent annotation measurement list. This one calls one of two different IJ scripts, and inserts variables for thresholds given by the user. Maybe these scripts will help:Ī quick intro that also includes a link to Pete’s intro:Ī more general script to handle finding brighter regions rather than darker regions In Preferences, there should be a field for “ImageJ plugins directory.” I have created a few different scripts that use the Analyze Particles… feature to get objects returned to QuPath through ImageJ, so that is definitely possible. You do need to add the plugins to QuPath’s version of ImageJ though. That may or may not be helpful to you!įor the ImageJ stuff, you can run quite a few plugins from ImageJ through QuPath, though I have not specifically tried that one (Weka segmentation). If I know I wanted separate files and was starting with a multiscene CZI, I would probably save the initial image at full quality, split the scenes, and then save with JPEGXR compression as the final step. Compressing it a second time makes things worse. 
So a 200MB scene might become 6GB, while still maintaining all of the resolution of a 200MB compressed image. The greater issue for me tends to be that the initial multiscene files might be JPEGXR compressed, and after performing operations on the image (like stitching the tiles together), the resulting image is full size. If over that limit, Zen (Blue) automatically creates it when you split scenes, write files, and look at the image (I think you need to open it). Zen will only create a pyramidal structure if the file size is over a certain limit, though I am not sure what that is. In the future we’d like to perform registration of multiple staining batches of the same sections that have been slide scanned so we can say perform 2 lots of 3 colour fluorescence staining and overlay them so we have six colours on the same image but I’m not sure if Qupath would manage this? I have played around to be able to export multiple TIFF files from Qupath so that’s the only way we can perform the analysis at the moment. Often we are quantifying multiple stromal markers and so need percentage area which seems to be more difficult to achieve with Qupath than with the analyze particles plugin in Image J. I’ve installed the Weka segmentation plugin on my Qupath software but the interface is very different and I’m struggling with it as I’d like to use it in the same way as Image J. We use Qupath and we think it handles these files brilliantly, however the issues we have are not being able to complete the same analysis in Qupath that we can perform in Image J.įor instance I use the Weka segmentation plugin in Image J to pull out areas of staining as we had some stained tissues sections using two colours brown and purple (plus hematoxylin) which didn’t seem to separate well using colour deconvolution. I also wrote a blog post showing how you can go from binary images showing detected regions back into QuPath here. 
There’s a video showing the general idea of exchanging data between QuPath and ImageJ here and some written documentation here. Alternatively, you could could use CellProfiler or some other software if you preferred for the batch processing. For example, you might save some TIFFs created in this way, run a Fiji macro/script to process them, and at some later date decide to write a QuPath script to bring all your ImageJ/Fiji ROIs back onto the whole slide. The properties are also maintained if you save the image as an ImageJ TIFF, which increases your processing options further. 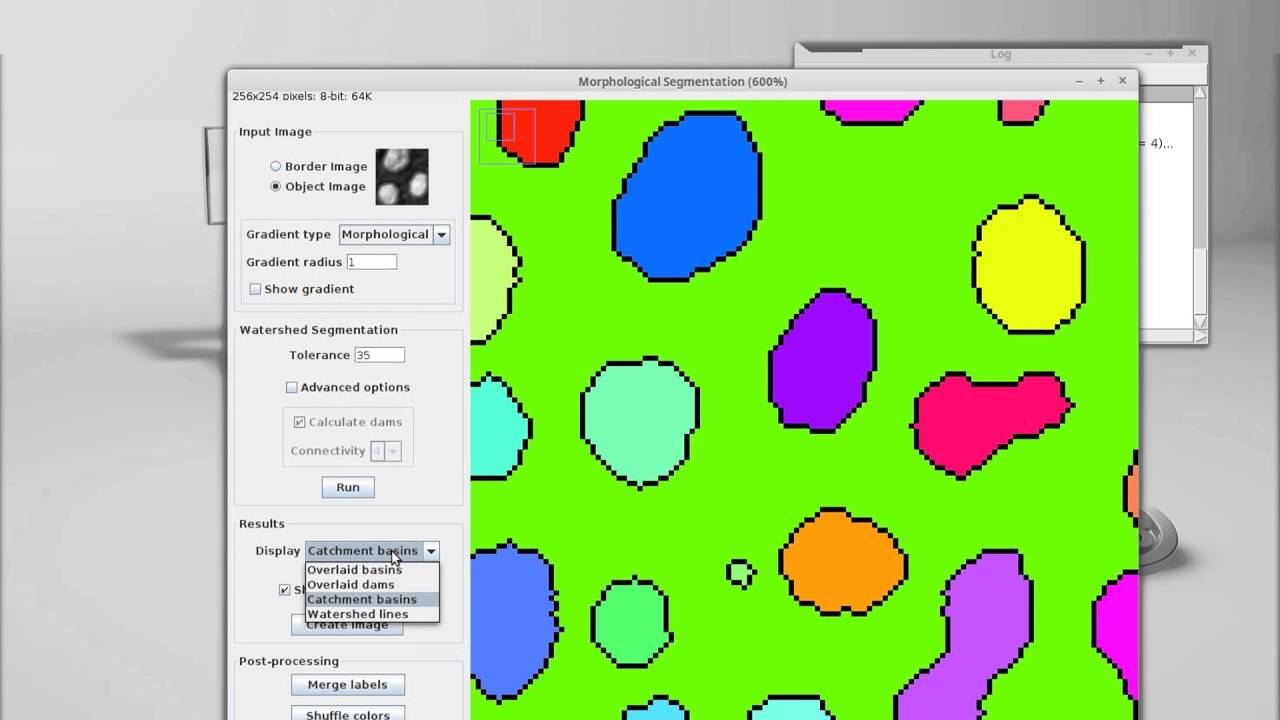
This means that if you create ROIs/overlays in ImageJ, you can optionally send them back to QuPath to view them in the correct location in the original whole slide image. The image sent to ImageJ should retain the channels and bit depth, and also have its properties set accordingly (e.g. Rather, you would draw around any region in QuPath and choose the option to send it to ImageJ - optionally scaling it down if you like. It also doesn’t have the conversion-to-RGB problem or any need to extract channels individually. Without meaning to push the QuPath thing too hard, assuming your image can be opened with QuPath + Bio-Formats, then that it is arguably easier than the Zen method.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
